Tools to compute functional diversity



Quantifying the relevance of intraspecific trait variabilty for functional diversity






Comparing alpha-, beta- and gamma- diversities across taxonomical, functional and phylogentic facets of biodiversity
Recently, use of the Rao quadratic entropy index has been advocated since it allows comparing the partitioning of various facets of diversity (e.g. taxonomic, phylogenetic and functional) within the same mathematical framework. However, if not well implemented, the Rao index can easily yield biologically meaningless results and lead into a mathematical labyrinth (JVS-Theseus.pdf). We propose a an R function (here or link beside) which performs the key computations solving these limitations using the Jost correction for the Rao index.

Functional diversity, i.e. extent of functional trait dissimilarities within and among ecological communities
For the macro to calculate functional diversity and trait overlap in Excel (by Jan Lepš) click here.
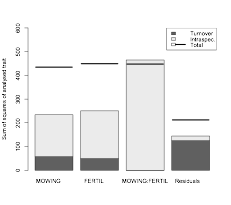
To what extent community trait composition across habitats is because of changes in species composition (i.e. species turnover) vs. intraspecific trait variability? The R function (AppendixEcography.zip) built by Petr Smilauer, allows to answer this question. The function provides nice graphs together with the complete analysis which can be carried out for any combination of categorial and quantitative predictors.



Community trait response to environment: disentangling species turnover vs. intraspecific trait variability effects
Intraspecific trait variability is a crucial, often neglected,
component of functional diversity. How to decompose the total functional diversity of a community into within-species vs. between-species effects? The R functions we propose are accompanied by a tutorial that explain how to implement them (figure beside or here).
A common requirement for computing most functional diversity indices is estimating trait differences between species. In this paper, using simulations we compared the two main approaches to compute trait differences: the Gower distance (using only species trait averages) and trait overlap (using intraspecific trait values). We discuss the major properties of these two approaches and provide guidelines for their applications. The paper will be published as part of a special issue on Journal of Vegetation Science on “Functional Diversity” in 2013. In the paper we also propose a new R function (“trova”, i.e. TRait OVerlAp), mainly done by Carlos Perez Carmona, which performs computations to estimate species trait dissimilarity with different types of data. Note that “trova” is also a nice cuban music played mainly with guitar (http://en.wikipedia.org/wiki/Trova) and in italian it means “to find”.
Trova, an integrated R function to compute different measures of trait dissimilarity between species

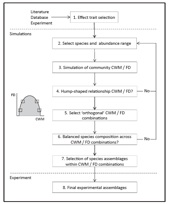
There is a growing consensus that the distribution of species trait values in a community can greatly determine ecosystem processes and services delivery. Two distinct components of community trait composition are hypothesized to chiefly affect ecosystem processes: (i) the average trait value of the species, quantified by community-weighted mean trait values (CWM; related to the mass ratio hypothesis) and (ii) the degree to which trait values differ between species in a community, quantified by different indices of functional diversity (FD; related to non-additive community effects). The interdependence between CWM and FD (CWM and FD generally showed a hump-shaped relationship i.e. at more extreme CWM values, communities can have only low FD values) poses a challenge on disentangling their relative importance using empirical data. We present a framework that allows designing manipulative experiments to decouple and assess the effects of these two community functional components on ecosystem processes an services. A R function (funziona) and a script to run such analyses is also given.
Funziona: a function to design experiments to disentangle the effects of mass ratio vs. complementarity hypotheses (CWM vs FD)